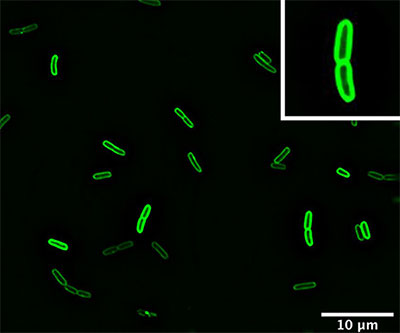
Microscopy image showing AzuC labeled with green fluorescent protein.
Credit: Storz Lab, NICHD
Small RNAs (sRNAs) help regulate gene expression—the degree to which genes are turned “on” or “off”—by interacting with messenger RNA (mRNA), the intermediary between DNA and protein. Although most sRNAs do not contain instructions for making a protein, some encode a small protein in addition to their regulatory role. The Storz Lab recently identified and characterized two such dual-function sRNAs.
The researchers found that Spot 42, a well-characterized sRNA that helps regulate carbohydrate metabolism in the bacterium Escherichia coli, surprisingly also encodes a tiny protein called SpfP. In the presence of the sugar glucose, E. coli’s preferred carbon source, Spot 42 blocks the expression of proteins involved in metabolizing non-preferred carbon sources like galactose. The NICHD scientists found that SpfP prevents bacterial growth in the presence of galactose. Further experiments revealed that SpfP binds to the CRP transcription factor (a protein that controls gene expression), blocking its ability to activate certain genes. This finding raises the question of how many other bacterial transcription factors are regulated by unidentified small proteins.
In a separate study, the Storz Lab characterized AzuCR, a dual-function sRNA from E. coli with competing regulatory functions. The scientists discovered that as a consequence of stimulating a key enzyme, AzuC protein reduces bacterial growth in glycerol. A segment of the sRNA itself, denoted AzuR, also base pairs with and blocks expression of two mRNAs. Interestingly, the researchers found that AzuC translation—the process by which RNA is converted to protein—interferes with repression of the mRNAs. Likewise, base pairing of AzuR with the mRNAs interferes with AzuC translation, demonstrating that the two regulatory functions of AzuCR compete.
Learn more about the Cell and Structural Biology Group: https://www.nichd.nih.gov/about/org/dir/affinity-groups/CSB
 BACK TO TOP
BACK TO TOP