Home
Most organisms have multiple forms of RNase H; two RNases H that are dissimilar in primary amino acid sequence yet have similar structural and, presumably, a common enzymatic mechanism. To address the cellular roles of these RNases H, we have examined the effects of eliminating RNases H in unicellular organisms such as bacteria and fungi. We have developed several mouse lines that are permitting us to see in detail the role for RNase H1 in replication of mitochondrial DNA. Patients with mutated RNase H1 exhibit typical muscular defects observed in other mitochondrial-related disease. Severe neuroinflammatory issues are seen in Aicardi-Goutieres Syndrome caused my defects in RNase H2. Major substrates for RNases H for RNases H are RNA/DNA hybrids that form during transcription. Failure to resolve these R-loops can have wide-spread effects that can lead to genome instability.
Click image to enlarge.
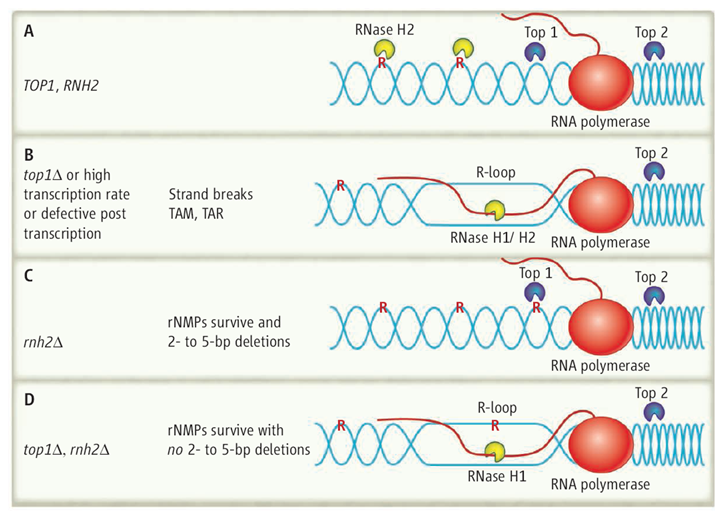
On top. (A) In yeast wild-type cells (TOP1, RNase H2), R-loop formation is prevented when Top2 relieves the positive supercoiling ahead of the transcribing RNA polymerase, and Top1 relieves the negative supercoiling behind. RNase H2 cleaves rNMPs incorporated into DNA, which leads to their replacement with dNMP. (B) In top1Δ mutants, RNA/DNA hybrids are formed and processed by RNase H1/H2, and rNMP removal is initiated by RNase H2. Mutations can result by TAM and TAR. (C) In rnh2Δ cells, Top1 will encounter and cleave rNMPs that have not been removed because RNase H2 is missing, which produces 2- to 5-bp deletions. (D) In the double mutant top1Δ rnh2Δ, RNA in R-loops may be cleaved by RNase H1, but rNMPs will remain in the DNA. DNA is blue, RNA is red, and single rNMPs in DNA are noted as red R.
 BACK TO TOP
BACK TO TOP