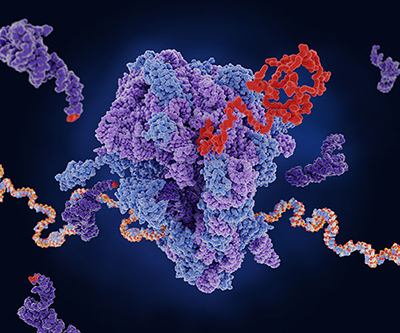
A ribosome (center) translates mRNA (multi-colored strand) into a protein (red). tRNA (dark purple) carries amino acids to add to the growing chain.
RNAs carry various instructions for cells. Messenger RNAs are translated into proteins, and non-coding RNAs, such as tRNAs, carry out important functions in a cell. Mutations in enzymes that modify tRNAs can lead to a variety of human diseases. About 10 years ago, the Maraia Lab developed an assay, based on a northern blot, to identify these types of mutations in tRNA modification enzymes (TME).
The method, called PHAM (i.e., positive hybridization in the absence of modification), has been used by several laboratories to produce molecular diagnostic criteria. It can identify modifications to both cytoplasmic and mitochondrial tRNAs. To date, PHAM has helped identify four founding cases of human diseases caused by TME mutations.
Among RNA, tRNAs are the most extensive post-transcriptionally modified, both in density (the percent of modified nucleotides) and diversity (different types of modifications). PHAM works by detecting the presence or absence of modifications on tRNAs to assess the extent of suspected or actual TME dysfunction. The assay focuses on modifications in specific areas of tRNAs called the anticodon loop and anticodon stem-loop. Modifications in these areas can influence features such as reading frame maintenance and translation efficiency, although anticodon loop modifications are especially important for proper deciphering of the large amount of redundant information in the genetic code.
In a recent publication, the Maraia Lab offers simple revisions to the method, such as customizing the wash temperature to each probe, that enhance the performance of PHAM. They also created a calculator to help researchers identify optimal conditions. These improvements help reduce variability in results and can lead to wider adoption of the method. Overall, PHAM is relatively easy assay to carry out in the laboratory. It’s considered a fast, reliable, and quantitative alternate to more complex, costly, and training-intensive methods, such as mass spectroscopy.
 BACK TO TOP
BACK TO TOP