Brain Browser
The Zebrafish Brain Browser is a tool for 3D visualization of transgene and gene expression patterns in 6 dpf larval zebrafish. At present, the database includes expression patterns for 264 transgenic, enhancer trap or immunostained larvae, each scanned at high resolution and registered to a single reference brain. The browser enables expression patterns to be compared at near single cell resolution, and the identification of transgenic lines that express Gal4 in almost any selected set of neurons. More information is available in Tabor (2019) .
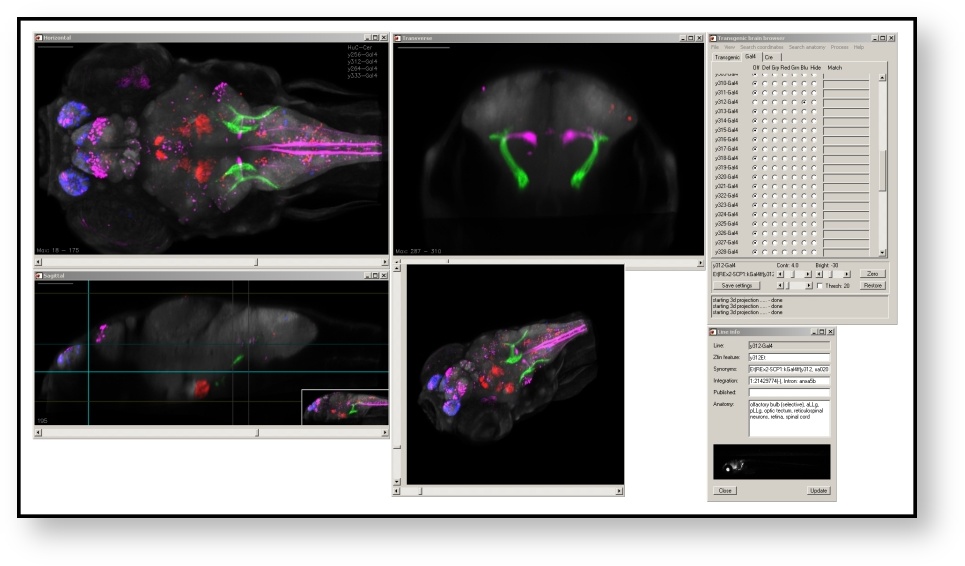
Access the Brain Browser via an online interface .
Alternatively, you can locally run a desktop version (via Zenodo ; 5.9 GB, ZBB2.0) that includes software, brain scans and anatomical regions (as described in Z-Brain , which we aligned to the Brain Browser using the method described in Marquart (2016) ). Contact us if you need an older version for any reason.
Both the online and local versions of the Browser include links to Zfin for ordering lines.
The Brain Browser was designed to make it simple to add additional data. To do this you need to:
- Confocal scan your transgenic reporter, crossed to the vGlut:DsRed transgenic line (or other suitable pattern): detailed imaging protocol here (PDF 653 KB).
- Register your confocal stack to the vGlut:DsRed reference brain we used, either:
a. (good) by using CMTK - instructions on installing and using CMTK for brain registration in this protocol (PDF 446 KB).
b. (better) by using ANTs , as described in Marquart (2016) . - Drag and drop the registered stack into the images directory of the browser.
A variety of transgenic lines can be used for registration, available here .
We can provide additional help with each of these steps. We welcome feedback on bugs in the software and suggestions for new features. If you'd like to contribute transgenic lines or confocal scans, that would be great too.
Virtual Reality Fish Brain
See the zebrafish brain in 3D, using our virtual reality Brain Browser app for google cardboard: brainbrowser.apk (APK 42 MB) (Android only for now).
- To navigate, toggle movement using the cardboard button.
- Switch on/off the visibility of elements using the panel at the button of the arena. Aim the cursor at a panel, and depress the cardboard button.
The VR Browser was developed by Damian Dalle Nogare (NICHD).

Additional resources
Other valuable tools for brain registration and analysis in zebrafish:
- Mapzebrain: Brain atlas for zebrafish, developed by the Baier lab : The Virtual Brain Explorer for Zebrafish, developed by the Driever lab.
- Z-Brain : A neuroanatomical reference atlas, developed by the Engert and Schier labs.
 BACK TO TOP
BACK TO TOP