Research
From the NICHD Annual Report:
In our tissue-sciences research, we strive to understand fundamental relationships between function and structure in living tissues. Specifically, we are interested in how tissue microstructure, hierarchical organization, composition, and material properties all work together to affect biological function or dysfunction. We investigate biological and physical model systems at various length and time scales, performing biophysical measurements and developing novel physical/mathematical models (including molecular dynamics [MD] and continuum models) to explain their functional properties and behavior. Experimentally, we often use water to probe tissue structure and function from nanometers to centimeters and from microseconds to lifetimes. Our armamentarium includes atomic force microscopy (AFM), small-angle X-ray scattering (SAXS), small-angle neutron scattering (SANS), static light scattering (SLS), dynamic light scattering (DLS), osmometry, and multi-dimensional nuclear magnetic resonance (NMR) relaxometry and diffusometry. A goal is to develop understanding and tools that can be translated from bench-based quantitative methodologies to the bedside to aid in diagnosis and therapy.
These activities dovetail with our basic and applied research in quantitative imaging, which is intended to generate measurements and maps of intrinsic physical quantities, including diffusivities, relaxivities, exchange rates, etc., rather than the qualitative ‘weighted’ MR images conventionally used in radiology. At a basic level, our work is directed toward making critical ‘invisible’ biological structures and processes ‘visible.’ Our quantitative imaging group uses their knowledge of physics, engineering, applied mathematics, imaging and computer sciences, as well as key insights gleaned from our tissue-sciences research, to discover and develop novel quantitative imaging biomarkers that can detect changes in tissue composition, microstructure, and/or microdynamics with high sensitivity and specificity. The ultimate translational goal is to assess normal and abnormal developmental trajectories, diagnose childhood diseases and disorders, and characterize degeneration and trauma (such as mild traumatic brain injury). Primarily, we use MRI as our imaging modality of choice because it is so well suited to many applications critical to the NICHD mission; it is non-invasive, non-ionizing, usually requires no exogenous contrast agents or dyes, and is generally deemed safe and effective for use with mothers, fetuses, and children in both clinical and research settings.
One of our technical translational goals has been to transform clinical MRI scanners into scientific instruments capable of producing reproducible, accurate, and precise imaging data with which to measure and map useful imaging biomarkers for various clinical applications, including single scans, longitudinal and multi-site studies, personalized medicine, and genotype/phenotype correlation studies, as well as for populating imaging databases with high-quality normative data. From a more basic perspective, another goal has been to apply our various MRI tools and methodologies to advance neuroscience, providing new methods to explore brain structure/function relationships, such as the human connectome.
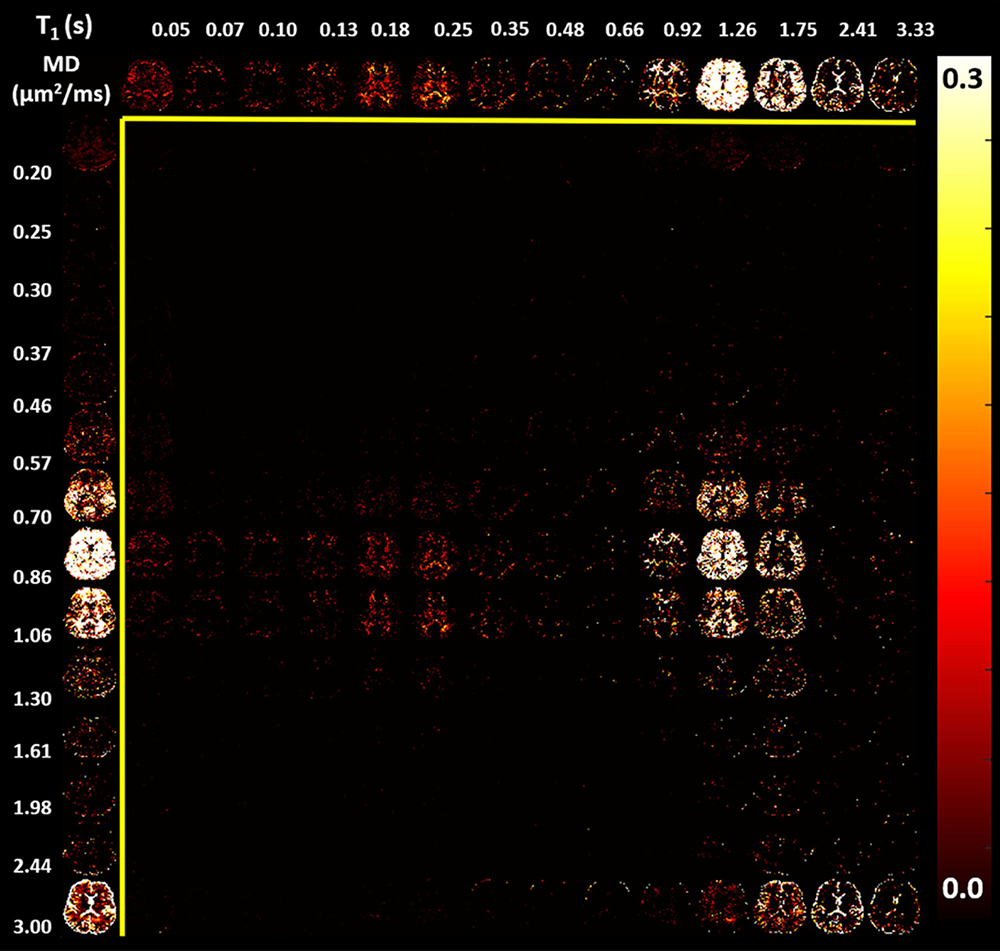
Figure 1. Maps of an estimated T1–MD (mean diffusivity) 2-D spectrum obtained in human brain in vivo in a clinically feasible 20-minute MRI scan
Click image to view.
Top row. The corresponding 1-D marginal distribution of T1 (nuclear spin relaxation time) values.
Left column. The corresponding 1-D marginal distribution of MD values. The high-dimensional relaxometry-diffusometry imaging approach allows us to identify and characterize the state of different and distinct water pools within each voxel, rather than report only their voxel-averaged values, which has traditionally been done. One can see various water compartments, such as high T1 and high MD, corresponding to cerebral spinal fluid; intermediate T1 and MD, corresponding to gray and white matter parenchymal tissue; and short T1 and intermediate MD, corresponding to white matter. This new method offers great promise for probing different tissue components occupying the voxel on a whole-brain basis to try to detect normal and abnormal development, degeneration, or trauma.
In vivo MRI histology
The most mature in vivo MRI histological technology that we invented, developed, and clinically translated is Diffusion Tensor MRI (DTI), by which we measure and map D, a diffusion tensor of water within an imaging volume. Information derived from this quantity includes white-matter fiber-tract orientation, the orientationally averaged mean apparent diffusion constant (mADC), and other intrinsic scalar (invariant) quantities. Such imaging parameters have been used by radiologists and neuroscientists as non-invasive quantitative histological ‘stains,’ MRI images that are obtained by probing endogenous tissue water in vivo without requiring any exogenous contrast agents or dyes. The mADC is the most widely used diffusion imaging parameter in neuroradiology to identify ischemic areas in the brain during acute stroke and to follow cancer patients’ responses to therapy. The measures of diffusion anisotropy we first proposed (e.g., the fractional anisotropy or FA) are also widely used to follow changes in normally and abnormally developing white matter and in many other clinical and neuroscience applications, such as brain white-matter visualization. Our group also pioneered the use of fiber direction–encoded color (DEC) maps to display the orientation of the main association, projection, and commissural white matter pathways in the brain. To assess anatomical connectivity among various cortical and deep-brain gray-matter areas, we also proposed and developed DTI ‘Streamline’ Tractography, which is used brain track white-matter fibers to help establish 'anatomical connectivity' and by neuroradiologists and neurosurgeons to plan surgical procedures so that they can spare 'eloquent' areas of the brain, advances that helped inspire several large federally-funded research initiatives, including the NIH Human Connectome Project (HCP) and, more recently, the NIH Brain Initiative.
More recently, we invented and developed a family of advanced in vivo diffusion MRI methods to measure fine-scale microstructural features of axons and fascicles (also known as ‘microstructure imaging’), which otherwise could only be assessed using laborious ex vivo histological or pathological methods. We have been developing efficient means for performing ‘k- and q-space MRI’ in the living human brain, such as ‘Mean Apparent Propagator’ (MAP) MRI, an approach that can detect subtle microstructural and architectural features in both gray and white matter at micron-scale resolution, several orders of magnitude smaller than the typical MRI voxel. MAP MRI also subsumes DTI, as well as providing a bevy of new in vivo quantitative ‘stains’ or biomarkers to measure and map. We also developed CHARMED MRI, which measures the average axon diameter (AAD), and AxCaliber MRI, which measures the axon-diameter distribution (ADD) along white-matter pathways, and we reported the first in vivo measurement of ADDs within the rodent corpus callosum. The ADD is functionally important, given that axon diameter is a critical determinant of conduction velocity and therefore the rate at which information travels along axon bundles, and helps determine the latencies or time delays between and among different brain areas. This led us to propose a novel MRI–based method to measure the ‘latency connectome,’ including a latency matrix that reports conduction times between different brain areas, and to develop a companion mathematical theory to explain the observed ADDs in different fascicles, suggesting that they represent a trade-off between maximizing information flow and minimizing metabolic demands. We developed novel multiple pulsed-field gradient (mPFG) methods and demonstrated their feasibility in vivo on conventional clinical MRI scanners as a further means to extract quantitative features in the central nervous system (CNS), such as the AAD and other features of cell size and shape.
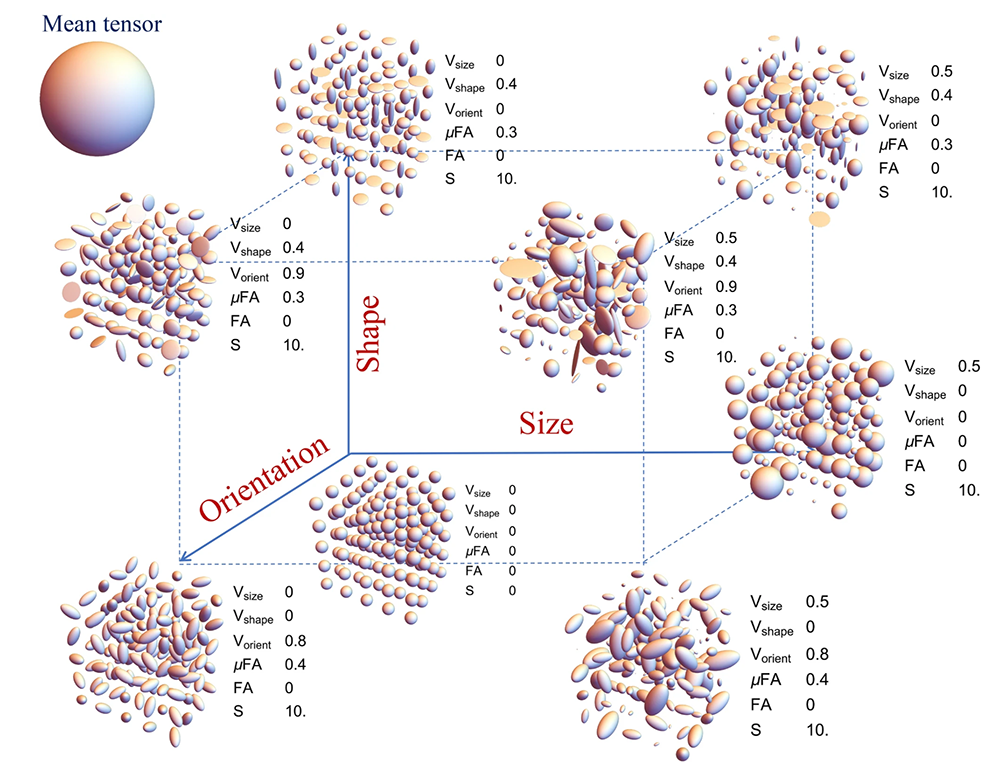
Figure 2. Diffusion Tensor Distribution (DTD) MRI resolves microscopic heterogeneity in brain tissue
Click image to view.
A key issue we addressed is that diffusion tensor MRI (DTI) can produce the same tensor estimates in different voxels that may have vastly different underlying tissue microstructures. In order to resolve this problem, we developed various new parametric quantitative imaging ‘stains’ to depict the shape, size, and orientation heterogeneity of the underlying distribution of diffusion tensors within a voxel reflecting different microstructural motifs. These are shown here for a macroscopically isotropic mean diffusion tensor (shown as a sphere in the upper left corner). The quantities are derived from a new imaging method we developed to measure or estimate a DTD in each voxel. Values of the various stains are shown as points in the 3D grid. Size axis refers to heterogeneity in the trace or mean diffusivity of the diffusion tensors; the shape axis refers to heterogeneity in the shapes of the diffusion tensors (i.e., oblate, prolate, spherical); and the orientation axis refers to the heterogeneity vs. coherence of the orientations of the diffusion tensors. Vsize, stain quantifying the variation in the size (i.e., trace) of the diffusion tensors; Vshape, stain quantifying the variation in the shape of diffusion tensors; Vorient, stain quantifying the orientation dispersion of diffusion tensors. The micro Fractional Anisotropy, micro FA, and macro FA obtained from DTI are also shown, as well as S, the overall entropy of the DTD, as a measure of tissue complexity.
Although gray matter appears featureless in DTI brain maps, its microstructure and architecture are rich and varied throughout the brain, not only along the brain's cortical surface, but also within and among its various cortical layers and within deep gray-matter regions. To target this tissue, we have been developing several non-invasive, in vivo methods to measure unique features of cortical gray-matter microstructure and architecture that are visible in electron microscopy (EM) applications but currently invisible in conventional MRI. One example is diffusion tensor distribution (DTD) MRI, in which we use our normal tensor-variate distribution to characterize heterogeneities in complex tissue structure. One of our long-term goals is to ‘parcellate’ or segment the cerebral cortex in vivo into its approximately 500 distinct cyto-architechtonic areas using non-invasive imaging methods. To this end, we are developing advanced MRI sequences to probe correlations among microscopic displacements of water molecules in the cortex as well as sophisticated mathematical models to infer distinguishing microstructural and morphological features of cortical gray matter. One promising avenue is to use multi-dimensional MRI relaxometry and diffusometry-based methods to study water mobility and exchange in gray and white matter, enabling us to probe individual water pools or compartments that are too small to observe using conventional MRI. We continue to work to translate these and other methods to the clinic to help identify changes in normal and abnormal development, as well as in inflammation and trauma.
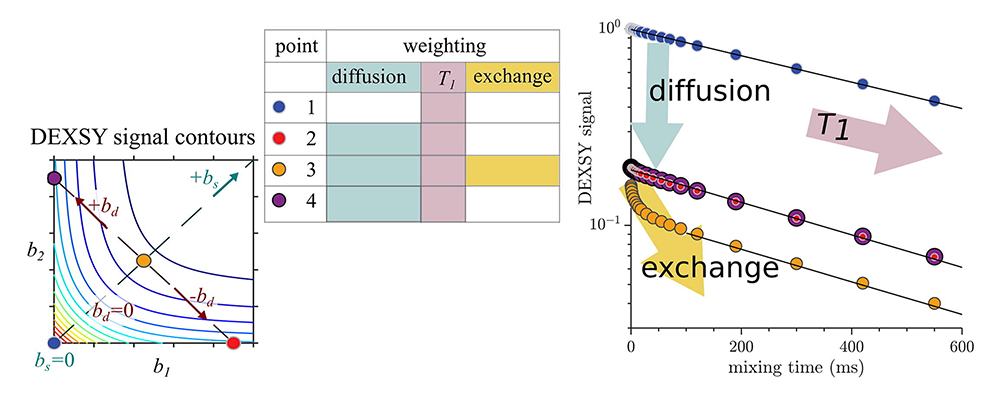
Figure 3. Deconflating effects of restriction and exchange in diffusion MR data
Click image to view.
We developed a novel method for acquiring diffusion exchange spectroscopy (DEXSY) MR data, and a novel use as a potential in vivo functional imaging (fMRI) tool, an innovation that required (a) vastly speeding the MR DEXSY signal acquisition and (b) being able to disambiguate the effects of two key processes previously conflated in the analysis of DEXSY data: restriction or confinement of water in pools or compartments, and exchange between different water compartments.
Quantitative MRI biomarker development for pediatric and fetal imaging applications
MRI is considered safer than X-ray–based methods, such as computed tomography (CT), for scanning fetuses, infants, and children. However, clinical MRI still lacks the quantitative character of CT. The scope of conventional MRI clinical applications is limited to revealing either gross morphological features or focal abnormalities. Clinical MRI also often lacks the biological specificity necessary for developing robust and reliable imaging ‘biomarkers.’ In particular, MRI assessment of normal brain development and developmental disorders has benefited greatly from the introduction of ‘quantitative’ clinical MRI techniques, with which one measures and maps meaningful intrinsic physical quantities or chemical variables that possess physical units and can be compared among different tissue regions. Quantitative MRI methods such as DTI also increase sensitivity, providing a basis for monitoring subtle changes that occur, e.g., during the progression or remission of disease, by comparing measurements in a single subject against normative values obtained from a healthy population. Quantitative MRI methods should continue to advance ‘precision imaging’ studies, in which MRI phenotypic and genotypic data can be meaningfully incorporated and used for improved diagnosis and prognosis assessments.
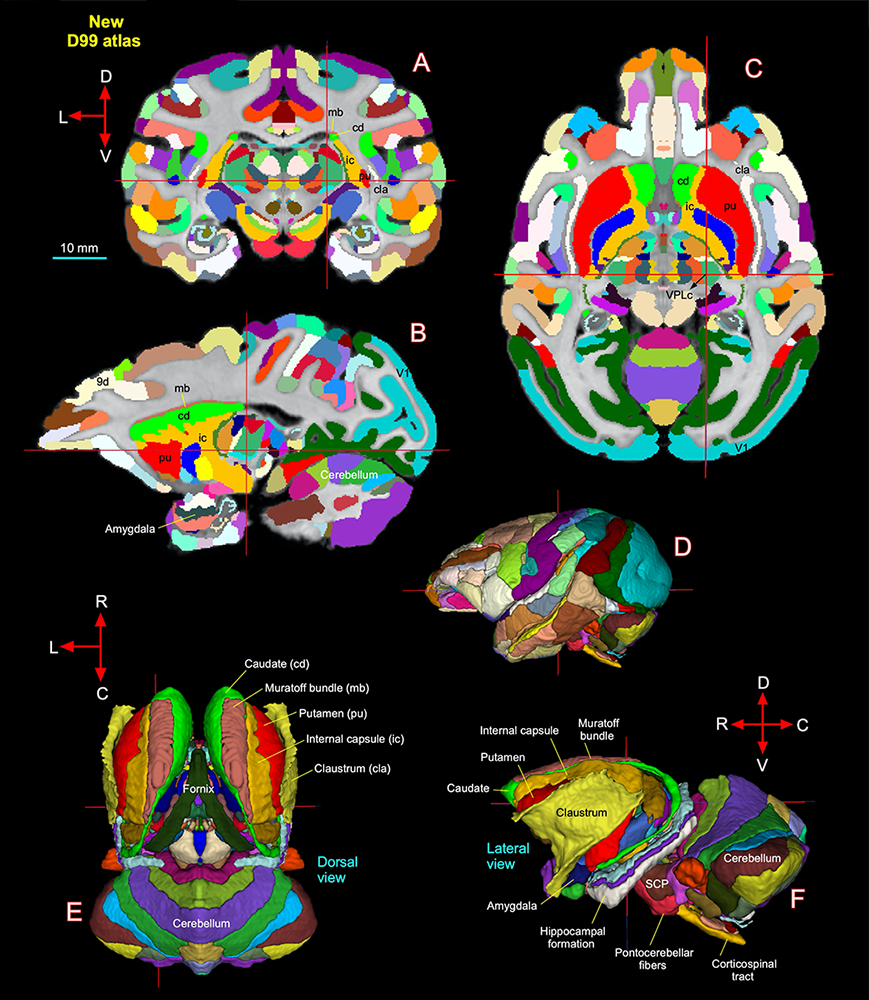
Figure 4. New macaque brain atlas uses Mean Apparent Propagator (MAP) MRI to improve delineation of different brain areas and structures
Click image to view.
A–C. New D99 digital macaque atlas (version 2.0) with combined cortical and subcortical segmentation overlaid on the coronal, sagittal, and horizontal D99 ex vivo MRI template, respectively. The cross-hairs in A–C show the same location of thalamic subregion VPLc (ventral posterior lateral caudal nucleus). The corresponding anteroposterior location of VPLc is also shown with the D99 rendered brain volume in D.
E–F. The spatial location of segmented subcortical regions shown on the dorsal and lateral views in 3D. The selected subcortical regions in E–F are also indicated with cortical areas in A–C. 9d, dorsal prefrontal area; SCP, superior cerebellar peduncle; V1, primary visual cortex. D, dorsal; V, ventral; R, rostral; C, caudal; L, lateral. Scale bar: 10 mm, applies to A–C only.
To advance our quantitative imaging activities, we developed numerical and statistical methods, including algorithms that generate a continuous, smooth approximation to the discrete, noisy, measured DTI field data so as to reduce noise and which allowed us to implement Streamline Tractography. We proposed a novel Gaussian distribution for the tensor-valued random variables that we use to design optimal DTI experiments and interpret their results. In tandem, we developed non-parametric empirical (e.g., Bootstrap) methods to determine the statistical distribution of DTI–derived quantities in order to study, e.g., the inherent variability and reliability of computed white-matter trajectories, enabling us to apply powerful hypothesis tests to assess the statistical significance of findings in a wide range of important biological and clinical applications that had been tested using ad hoc statistical methods. We are also developing novel methods to register different brain volumes and to generate group-average DTI data or atlases from various subject populations based on the Kullback-Leibler divergence.
Previously, we carried out clinical studies that utilize novel quantitative MRI acquisition and analysis methods and whose aim is to improve accuracy and reproducibility of diagnosis and to detect and follow normal and abnormal development. One early example is the NIH Study of Normal Brain Development, jointly sponsored by the NICHD, NIMH, NINDS, and NIDA, which was initiated in 1998 and intended to advance our understanding of normal brain development in typical healthy children and adolescents. The Brain Development Cooperative Group 

Traumatic Brain Injury (TBI) represents a significant public health challenge for our pediatric population, but also for our young men and women who serve in the military. Our involvement in TBI research, particularly in detecting mild TBI (mTBI), has continued to expand through partnerships with various Department of Defense (DoD) entities. Diffusion MRI (dMRI) provides essential information to aid in the assessment of TBI, but conventional dMRI methods have lacked sufficient specificity. To improve the accuracy and reproducibility of MAP–MRI findings, we developed a data–processing pipeline, and, in collaboration with scientists at the DoD Center for Neuroscience and Regenerative Medicine (CNRM), performed the first normative MAP–MRI studies, and applied this new and powerful method to detect tissue damage in brains of individuals who have suffered TBI, extending our NICHD TORTOISE pipeline to be able to analyze MAP–MRI data. We are now employing multi-dimensional MRI relaxometry-diffusometry methods to study the etiology of various types of TBI, in collaboration with the USUHS Neuropathology Research Division and under the auspices of the CNRM, and to improve the correlation and integration of neuropathology and neuroradiological imaging data to speed the deployment of new MRI methods to assess various forms of TBI. We have also partnered with CNRM to study ways to measure very slow flows that occur during glymphatic transport, a mechanism the brain uses to wash away harmful macromolecules, just as the lymphatic system uses in other organs. With our partners at the University of Arizona, this research is providing experimental data to enable us migrate these imaging approaches to the clinical, to be able to assess normal and pathological glymphatic transport in vivo.
We are also collaborating with Sara Inati, who studies focal epilepsy, a devastating disorder that is difficult to detect using conventional neuroradiological methods. We are developing and testing various new MRI–based methods that we believe may reveal pathological microstructural features and changes in architectural organization of the brain in this disorder, for example, in cortical dysplasia, to improve localization and assessment of cortical lesions.
We have been partnering with our NICHD colleagues Roberto Romero and Mark Haacke to develop novel fetal MR imaging applications. Currently, it is challenging to measure quantitative imaging biomarkers in utero, particularly diffusion MRI–based biomarkers, owing to large-scale fetal and maternal motion during the scanning session, and to the difficulty in acquiring image volumes with sufficient coverage, quality, and spatial resolution in a clinically feasible amount of time. Our lab has been developing novel approaches to address these critical issues.
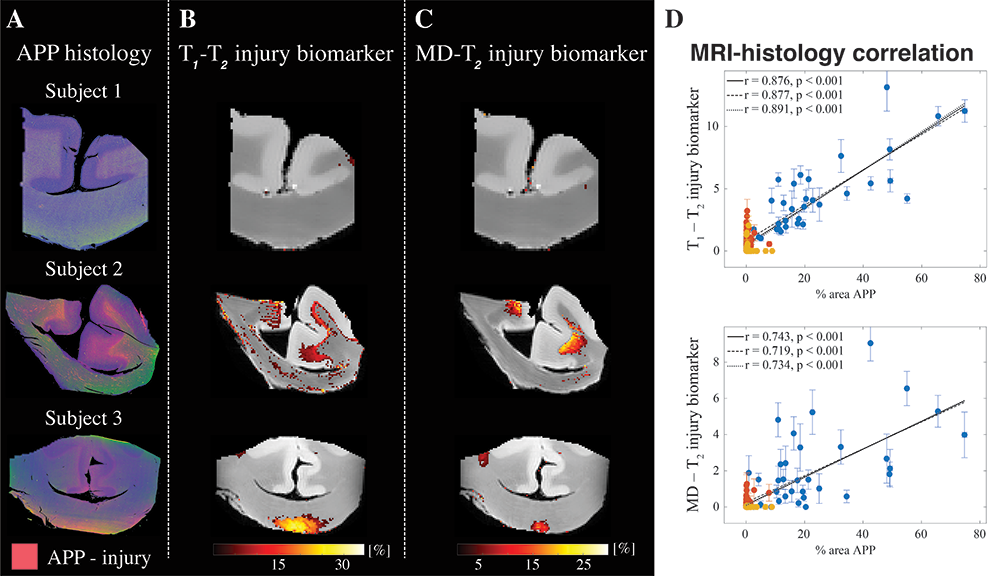
Figure 5. Potential MRI biomarker for TBI tested in a neuropathology brain specimen
Click image to view.
The figure suggests that MRI can now be used to detect sequelae of Traumatic Axonal Injury in neuropathological brain tissue specimens. Histological images juxtaposed with multidimensional MRI–derived injury biomarker maps of three representative TBI (traumatic axonal injury) cases (subjects 1, 2, and 3 are control and two TBI cases, respectively).
A. Deconvolved histological amyloid precursor protein (APP) images (co-registered with MR images) are shown on the left panel, red = APP.
B. T1–T2 and C Mean Diffusivity (MD)–T2 multidimensional injury maps show absence of significant injury, substantial injury along the white-gray matter interface, and substantial injury at the base of the corpus callosum, for subjects 1 to 3, respectively. All multidimensional injury maps were thresholded at 10% of the maximal intensity and overlaid on grayscale proton-density images.
D. APP density (% area) from 132 tissue regions in 11 different cases, consisting of 4 APP–positive regions from each TBI case (total of 32, blue dots), 4 to 6 normal-appearing white matter (WM) regions from all cases (total of 56, red dots), and 4 cortical grey matter (GM) regions from all cases (total of 44, yellow dots), and the corresponding MR parameter correlations. Individual data points represent the mean ROI value from each post-mortem tissue sample. Scatterplots of the mean (with 95% confidence interval error bars) % area APP and T1-T2 (top) or MD-T2 (bottom).
Biopolymer physics: water-ion-biopolymer interactions
Remarkably, despite their crucial role, little is known about the physical underpinnings of water-ion-biopolymer interactions, particularly in the physiological ionic strength regime. To determine the effect of ions on the structure and dynamics of key biopolymers, we developed a multi-scale experimental framework by combining macroscopic techniques (osmotic swelling-pressure measurements and mechanical measurements) with high-resolution scattering methods (e.g., SANS and SAXS) that probe the structure and interactions over a broad range of length and time scales. Macroscopic swelling-pressure measurements provide information on the overall thermodynamic response of the system, while SANS and SAXS allow us to investigate biopolymers at molecular and supramolecular length scales and to quantify the effect of changes in the environment (e.g., ion concentration, ion valance, pH, temperature) on the structure and interactions among biopolymers, water, and ions. Studies carried out on well defined model systems that mimic essential features of tissues provide important insights that cannot be obtained from experimental studies made on biological systems themselves. Mathematical models based on well established polymer-physics concepts and, more recently, molecular dynamics (MD) simulation approaches make it possible to design experiments, to help us quantify and explain aspects of tissue behavior and thus the underlying molecular and macroscopic mechanisms that govern key aspects of a tissue's normal functional properties.
We produced numerous novel biomimetic MRI phantoms, including diffusion MRI phantoms, which we use to calibrate scanners, specifically to assure the quality and fidelity of the imaging data in single-subject, longitudinal, and multi-site studies. For instance, our U.S. Patent for a ‘Phantom for diffusion MRI imaging’ is now enabling quantitative diffusion MRI studies to be performed at many different sites. Our colleagues at NIST Boulder, Colorado, have incorporated our polyvinylpyrrolidone (PVP) polymer into their own diffusion MRI NIST standard. We also developed a variety of NMR and MRI phantoms, such as 3-D printed polymer phantoms, that possess various salient features of cell or tissue systems, providing data with which to test the validity of our models and experimental designs, for instance in the testing of methods like our ‘AxCaliber MRI,’ which measures axon diameter distributions (ADD) in white matter pathways in vivo.
Measuring and mapping functional properties of extracellular matrix (ECM)
We study interactions among the main extracellular matrix (ECM) components, often using cartilage as a model system because it is aneural and avascular. In cartilage ECM, collagen (type II) is organized into fiber bundles that form a network that entraps the major proteoglycan (PG), a bottlebrush-shaped aggrecan. The biomechanical behavior of cartilage and other ECMs reflects their molecular composition and microstructure, which change during development, disease, degeneration, and aging. To determine tissue structure/function relationships, we measure various physical/chemical properties of ECM tissues and tissue analogs at different length and time scales, using a variety of complementary static and dynamic experimental techniques, e.g., osmometry, SANS, SAXS, neutron spin-echo (NSE), SLS, DLS, and AFM. Understanding the physical and chemical mechanisms affecting cartilage swelling (hydration) is essential to predicting its load-bearing ability, which is mainly governed by osmotic and electrostatic forces. To quantify the effect of hydration on cartilage properties, we previously developed a novel tissue micro-osmometer to perform precise and rapid measurements on small tissue samples (less than 1 microgram) as a function of the equilibrium water activity (vapor pressure). We also make osmotic pressure measurements to determine how the individual components of cartilage ECM (e.g., aggrecan and collagen) contribute to the total load-bearing capacity of the tissue. We also demonstrated that aggrecan-hyaluronic aggregates self-assemble into microgels, contributing to improved dimensional stability of the tissue and its lubricating ability. We also found that aggrecan is highly insensitive to changes in the ionic environment, particularly to calcium ions, which is critical for maintaining the tissue's mechanical integrity and allowing aggrecan to serve as a calcium ion reservoir in cartilage and bone.
To model cartilage ECM, we invented and developed a new biomimetic composite material consisting of polyacrylic acid (PAA) microgel particles dispersed and embedded within a polyvinyl alcohol (PVA) gel matrix. PAA mimics the proteoglycan (i.e., hyaluronic-aggrecan complexes), while PVA mimics the fibrous collagen network entrapping them. Remarkably, the PVA/PAA biomimetic model system reproduces not only the shape of the cartilage swelling pressure curves, but also the numerical stiffness values reported for healthy and osteoarthritic human cartilage samples. Studies on these model composite hydrogels should continue to yield invaluable insights into the effects of various macromolecular factors (matrix stiffness, swelling pressure, fixed-charge density, etc.) on the tissue's macroscopic mechanical/swelling properties, and ultimately its remarkable load-bearing and lubricating abilities, and their loss in various diseases and disorders, including osteoarthritis.
We are now attempting to translate our understanding of the structure-function relationships of ECM components to develop and design novel non-invasive MRI methods, with the aim of inferring ECM composition, patency, and functional properties in vivo. Our goal is to use MRI for early diagnosis of diseases of cartilage and other tissue and organs to follow normal and abnormal ECM development, which entails making ‘invisible’ components of ECM, (e.g., collagen and PGs) ‘visible’ to predict functional properties of the composite tissue, such as its load-bearing ability. One major obstacle is that protons bound to immobile species (e.g., collagen) are largely invisible with conventional MRI methods. However, magnetization exchange (MEX) MRI (as well as other related methods) make it possible to detect the bound protons indirectly by transferring their magnetization to the abundant free water protons surrounding them. It also enables us to estimate collagen content in tissue quantitatively. In pilot studies with Uzi Eliav (deceased) and Ed Mertz, we applied the new MEX MRI method to determine the concentration and distribution of the main macromolecular constituents in bovine femoral-head cartilage samples. The results were qualitatively consistent with those obtained by histological techniques, such as high-definition infrared (HDIRI) spectroscopic imaging. Our novel approach has the potential to map tissue structure and functional properties in vivo and non-invasively. We are now developing molecular dynamics (MD)–based models of cartilage and cartilage ECM analogs in order to interpret our experimental findings, develop and test novel hypotheses, and predict the behavior of our model system under different experimental conditions.
We also recently began using several novel one-sided NMR methodologies to study water relaxation, diffusion, and exchange behaviors in ECM as a means to characterize its critical functional properties. Velencia Witherspoon has been using these approaches to study the organization and structure of fascia. Our specialized NMR scanner/profilers are ideally suited to these tasks, as they can probe layered media, such as cartilage and fascia, using ultra-thin slices, almost as thin as a confocal microscope provides.
Additional Funding
- “Development of Bench and Pre-Clinical MRI Methods to Assess Glymphatic Clearance in the Living Brain.” 308811-4.01-60855, (CNRM-89-9237), which is under the joint auspices of the NIH, DoD, CNRM, and USUHS.
- “In vivo Brain Network Latency Mapping.” NIH BRAIN Initiative grant 1-R24-MH-109068-01.
- “Connectome 2.0: Developing the next generation human MRI scanner for bridging studies of the micro-, meso- and macro-connectome.” NIH BRAIN Initiative-funded 1U01EB026996-01.
- “Neuroradiology/Neuropathology Correlation/Integration Core.” 309698-4.01-65310, (CNRM-89-9921)
 BACK TO TOP
BACK TO TOP