Findings in cell culture suggest potential strategy for developing COVID-19 treatments
Credit: NIAID
Like all viruses, SARS-CoV-2, the virus that causes COVID-19, relies on the cells it infects to reproduce. The host cell provides the components and machinery to build new infectious viral particles. A new study sheds light on the importance of certain modifications made by the host cell to the SARS-CoV-2 spike protein, which the virus uses to enter human cells. These modifications, known as S-acylation, appear to be critical for the virus’ ability to infect cells. The findings suggest that blocking S-acylation could serve as a potential strategy for developing COVID-19 treatments.
The study was led by Anirban Banerjee, Ph.D., and Robbins Puthenveetil, Ph.D., of the Section on Structural and Chemical Biology of Membrane Proteins at NIH’s Eunice Kennedy Shriver National Institute of Child Health and Human Development (NICHD). NICHD researchers and their colleagues at NIH’s National Cancer Institute report the findings in the Journal of Biological Chemistry.
Background
While scientists have learned a great deal about SARS-CoV-2 since the virus emerged in 2019, many aspects of its biology remain poorly understood. Better understanding of the factors that drive replication and spread of SARS-CoV-2 can help them identify new targets for potential COVID-19 treatments.
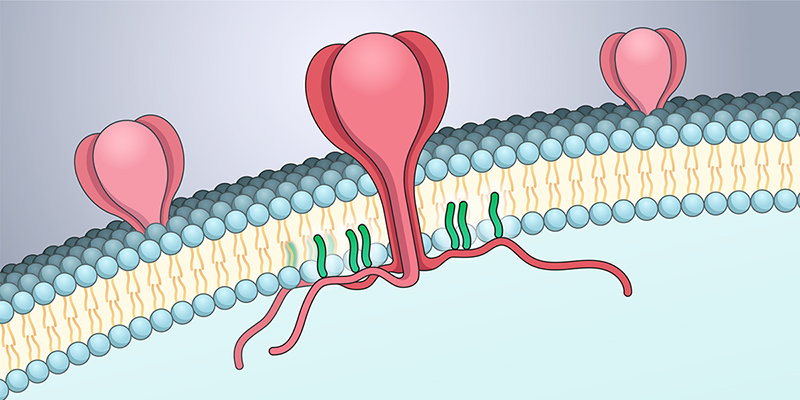
Credit: Nichole Swan, NICHD
SARS-CoV-2 uses its spike protein to enter human cells and start viral replication. Most of the spike protein sticks out from the surface of the virus, allowing it to recognize and bind to proteins on the surface of the human cells that the virus infects. While scientists have extensively studied the external part of the SARS-CoV-2 spike, little is known about the small portion of spike that resides internally within the virus. In the current study, researchers focused on this portion of the protein.
S-acylation, in which fatty acids are attached to proteins, can be crucial for a protein’s function. S-acylation of viral proteins relies on enzymes—proteins that facilitate chemical reactions—from the host cell.
S-acylation can occur only at certain sites on a protein. The interior portion of the SARS-CoV-2 spike protein harbors many possible sites for S-acylation. The NICHD-led team sought to determine the extent to which S-acylation of SARS-CoV-2 spike occurs and its potential impact on the virus’ ability to infect human cells.
Results
The researchers first looked at whether the SARS-CoV-2 spike undergoes S-acylation in human cells. Experiments in cell culture revealed that S-acylation indeed occurs and appears to be concentrated at three sites on the interior portion of spike. They then prepared a set of pseudoviruses—disabled viruses bearing the SARS-CoV-2 spike and spike variants engineered to lack S-acylation sites. They found that removing S-acylation sites greatly reduced or blocked the ability of the pseudoviruses to infect human cells in culture.
Next, the scientists sought to determine which human enzymes may be responsible for S-acylation of the SARS-CoV-2 spike. In human cells, the zDHHC family of 23 membrane-embedded enzymes carries out S-acylation of both human and viral proteins. A series of cell-culture experiments revealed seven zDHHC enzymes that potentially are involved in S-acylation of the SARS-CoV-2 spike.
Finally, the researchers developed a cell-free system that recapitulates S-acylation of the interior portion of the SARS-CoV-2 spike by human zDHHC enzymes. Such a system will make it easier and quicker to conduct experiments to further investigate the detailed mechanisms of SARS-CoV-2 spike S-acylation and to identify molecules that could block this process.
Significance
The results illuminate a poorly understood aspect of SARS-CoV-2 biology. They also suggest that blocking S-acylation of the SARS-CoV-2 spike protein could serve as a potential COVID-19 treatment strategy.
Next Steps
The authors note that further research is needed to more thoroughly understand S-acylation of the SARS-CoV-2 spike and its impact on the virus. The cell-free experimental system they developed, together with recently obtained atomic-level pictures of two human zDHHC enzymes, will help researchers identify molecules that can block S-acylation of the SARS-CoV-2 spike and other viral proteins.
Other NICHD co-authors on the paper include Elliot Murphy, Ph.D.; Liam Healy; and Eric Christenson, Ph.D.
Reference
Puthenveetil R et al. S-acylation of SARS-CoV-2 spike protein: mechanistic dissection, in vitro reconstitution and role in viral infectivity . Journal of Biological Chemistry DOI: 10.1016/j.jbc.2021.101112 (2021)
 BACK TO TOP
BACK TO TOP