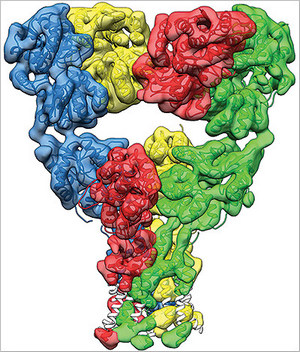
Scientists at the National Institutes of Health have created high-resolution images of the glutamate receptor, a protein that plays a key role in nerve signaling. The advance, published online in the journal Nature on August 3, 2014, opens a new window to study protein interactions in cell membranes in exquisite detail.
All cells have protein-based receptors on their surface, which act as specific docking sites for key chemicals that trigger a cellular reaction. Defects in these cell receptors are seen in diseases as far-ranging as cancer, Alzheimer's disease, and schizophrenia. Thus, revealing the shape of these cell receptors may enable scientists to craft therapies that target them.
The research team was led by Sriram Subramaniam, Ph.D., National Cancer Institute (NCI), in collaboration with colleagues at NIH, including Mark Mayer, Ph.D., Eunice Kennedy Shriver National Institute of Child Health and Human Development (NICHD), and at FEI, a company in Hillsboro, Oregon, that develops electron microscope technology.
The scientists used an imaging technique called cryo-electron microscopy (cryo-EM), an emerging tool for obtaining protein structures in various states. Cryo-EM is a more versatile approach to obtain protein structures than the commonly used method of X-ray crystallography, a process that requires scientists to force the protein to crystallize in a fixed shape.
Glutamate receptors are located in nerve cell membranes and are important for many functions in the nervous system, including memory. Their dysfunction has been implicated in neurodegenerative and psychiatric disorders, including Parkinson's disease and depression, and in some types of cancer. The glutamate receptor serves as a channel to allow certain atoms and molecules, called ions, into the nerve cell, which induces nerves to send signals..
"Ions are the currency of how brain cells talk to each other," said Mayer, an expert in the study of glutamate receptors.
How the glutamate receptor changes shape — to an open state to allow the transit of ions and then to a closed state again — is a question of great interest to scientists trying to understand how the brain works. The power of the cryo-EM method used by the NIH scientists is that they could capture the glutamate receptor in these different states and image them to derive the molecular movements involved.
What was revealed was a molecular machine with the precision of a fine Swiss watch, said Subramaniam, an expert in imaging. Obtaining these structural snapshots of the glutamate receptor in resting, open and desensitized states allowed visualization of the remarkable sequence of steps that enable control of ion flow across the cell membrane by these molecules.
"Not in a million years would I have dreamed that a cell receptor would work this way to open and close the gate for ion flow," Subramaniam said. "It is really satisfying to be able to decipher the inner workings of such an elegant molecular machine."
Understanding how the ion channels operate could lead to the creation of medications that inhibit or enhance these receptor motions. "Seeing the actual receptor in high resolution is half the battle," Mayer said.
The imaging feat builds upon other recent advances in cryo-EM pioneered by Subramaniam and his team, who reported in the Proceedings of the National Academy of Sciences on July 28, 2014, that the structure of the enzyme beta-galactosidase could be determined at a resolution of 3.2 angstroms, or 0.32 nanometers, using cryo-EM. (For reference, the diameter of a carbon atom is 1.7 angstroms.)
At these types of resolutions, the arrangement of individual amino acids in a protein can begin to be determined without the need to first get the proteins to form a crystalline lattice, as is done in crystallography, a step that doesn't always work for many classes of proteins.
"We are now poised to analyze structures of a wide variety of biologically and medically relevant multi-protein complexes and membrane protein assemblies, which have historically represented the most challenging frontier in structural biology," said Subramaniam.
Reference
Joel R. Meyerson (NCI), Janesh Kumar (NICHD), Sagar Chittori (NICHD), Prashant Rao (NCI), Jason Pierson (FEI Company), Alberto Bartesaghi (NCI), Mark L. Mayer (NICHD), and Sriram Subramaniam (NCI). Structural mechanism of glutamate receptor activation and desensitization. Nature. August 3, 2014. DOI: 10.1038/nature13603.
###
About the Eunice Kennedy Shriver National Institute of Child Health and Human Development (NICHD): The NICHD sponsors research on development, before and after birth; maternal, child, and family health; reproductive biology and population issues; and medical rehabilitation. For more information, visit the Institute's website at http://www.nichd.nih.gov.
About the National Institutes of Health (NIH): NIH, the nation's medical research agency, includes 27 Institutes and Centers and is a component of the U.S. Department of Health and Human Services. NIH is the primary federal agency conducting and supporting basic, clinical, and translational medical research, and is investigating the causes, treatments, and cures for both common and rare diseases. For more information about NIH and its programs, visit www.nih.gov.

 BACK TO TOP
BACK TO TOP